Research
Protein engineering
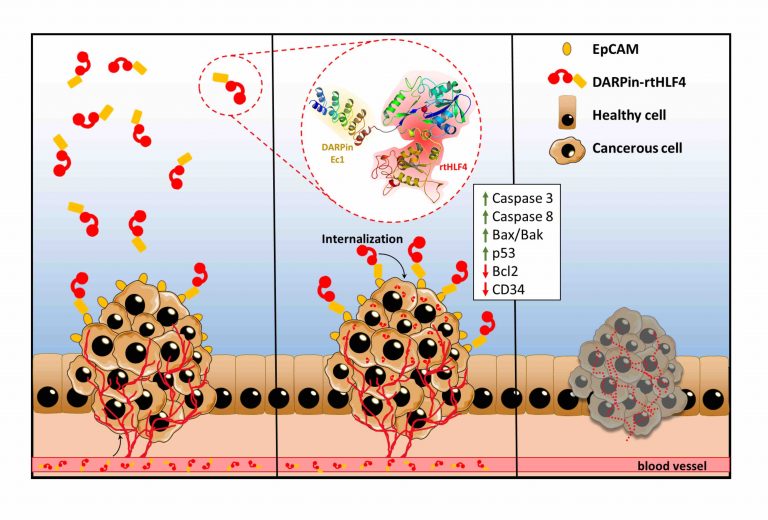
Protein-based therapeutics are considered a safer alternative to conventional drug therapies due to its high specificity and efficacy. However, such proteins generally have poor absorption and stability issues, and require invasive procedures while administration. Our study aims to increase the bioactivity of these proteins in the host body by increasing the specificity and efficacy by at least two to three orders of magnitude. First, we develop protein-tags for efficient protein therapeutics across the intestinal barrier. Second, we optimize therapeutic proteins to survive the harsh gastro-intestinal environment. Third, increase protein target specificity towards specific ailments. Our study aims to target two main ailments, cancer and infection. As a proof-of-concept, the study specifically looks into targeting pancreatic and breast cancer, and infiltrative microbial infections such as Helicobacter pylori and Fusobacterium nucleatum.
The development of these proteins has a two-pronged function. First, the direct use of these therapeutic protein as a bench-to-bedside therapy. Second, as an effector tool for sustainable delivery of therapeutic proteins using gut-residing probiotic and commensal microbes.
Microbiome
The microbiome is the collection of all microbes and their genes existing within a specific niche. The genes in the human microbiome outnumber the genes in our own genome by about 100 folds. The microbiome interacts with the host in various manners. As such, the human microbiome can be regarded as a human organ from a physiological standpoint, and its homeostasis is crucial to our health. Our study focuses on the bone, gut and lung microbiomes.
Bone microbiome
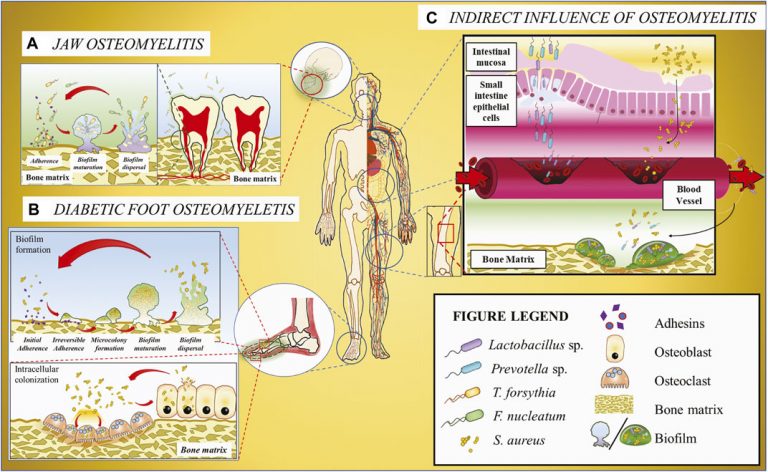
Although bone tissues of healthy humans remain considered sterile to date, the physiological structure of bones (including compact and cancellous bones) is suggestive of its susceptibility to exterior microbes, mainly through the blood. Interestingly, blood was also considered aseptic traditionally but has been acknowledged as septic recently, although the blood-borne microbes do not cause significant symptoms in healthy people. We found that microbes exist at low abundance in the bone tissues and these bone-derived microbial populations are dependent on the host species, age and health state.
Collaborating with Guangdong Second Provincial General Hospital, we are looking into how these microbial population impacts bone health by investigating the regulatory symbiotic function of these microbes on the osteoblast and osteoclast activities, and the onset of bone-related diseases such as osteomyelitis, osteoporosis and osteoarthritis.
The development of this study will allow a better understanding on the establishment of the bone microbiome, thus allowing long-term treatment in improving the host bone health by regulating the bone microbiota. Additionally, the characterization of bone commensal microbes provides a suitable chassis for the development of future engineered microbes for the treatment of bone fractures and diseases.
Gut microbiome
More than a trillion microbes live within our gastrointestinal tract and are responsible for vital metabolic, immune and nutritional functions. Dysbiosis is associated with many common diseases and is a target for therapy. Our study is focusing on kid inflammatory bowel disease (IBD) through collaborating with Shenzhen Children’s Hospital and Hong Kong Polytechnic University. IBD is a broad term that describes conditions characterized by chronic inflammations of the gastrointestinal tract.
By sequencing and analyzing the gut microbiome of IBD kids and the healthy controls, we will get information of the dysbiosis in IBD kids’ gut, and further recover the gut microbiome homeostasis by appropriate administration of prebiotics and probiotics.
Lung microbiome
Tuberculosis (TB) is a disease caused by a bacterium called Mycobacterium tuberculosis, which can spread from person to person through the air. TB is an opportunistic infection, which occurs more often or is more severe in people with weakened immune systems than in people with healthy immune systems. HIV weakens the host immune system, increasing the risk of TB in people with HIV. By collaborating with the Third People’s Hospital of Shenzhen, we are focusing on the investigation of the lung microbiomes of patients (including TB-infected, HIV-infected, and HIV/TB-coinfected) and healthy controls.
Data storage
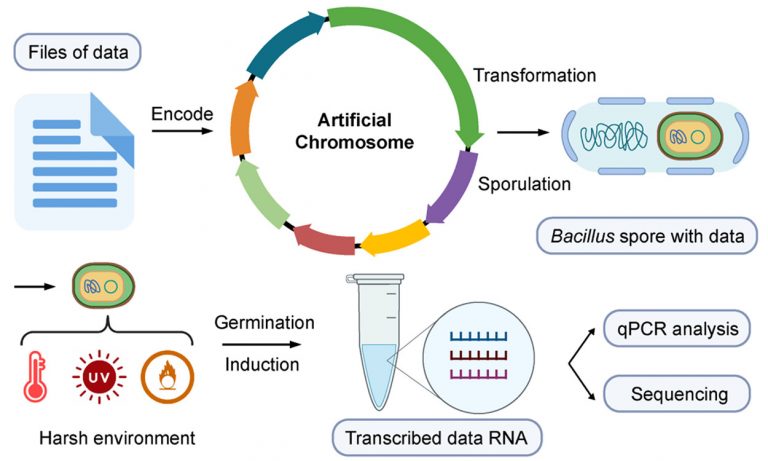
DNA data storage technology may supersede conventional chip or magnetic data storage medium, providing long-term stability, high density, and sustainable storage. Due to its error-correcting capability, DNA data stored in living organisms exhibits high fidelity in information replication.
Our team developed a Bacillus chassis integrated with an inducible artificially assembled bacterial chromosome to facilitate random data access. The sporulated Bacillus harboring DNA information could be used for long-term storage, where viability assays of spores were subjected to harsh environmental stresses to evaluate the data storage stability. The data accuracy remained above 99% after high temperature and oxidative stress treatment, whereas UV irradiation treatment provided above 96% accuracy. The developed Bacillus chassis and artificial chromosome facilitate the long-term storage of larger datum volume by using other DNA digital encoding and decoding programs.
Cyclic di-GMP
Bacteria have evolved a flexible self-protection mechanism to switch between motile (planktonic) state and sessile lifestyle (biofilm state). This switch process is regulated by the intracellular second messenger bis-(3′-5′)-cyclic dimeric guanosine monophosphate (c-di-GMP). However, closer understanding of the cellular processes resulting from the upregulation of c-di-GMP has not been fully revealed.
Our team investigates the role of cellular c-di-GMP in promoting planktonic cell growth in Escherichia coli K12 and Pseudomonas aeruginosa PAO1. We further identified key bacterial response genes linked to the upregulation of c-di-GMP that influences the observed phenotypic changes. This research could provide a glimpse on how the bacterial behavior evades potential threats through adapting bacterial phenotypic changes via c-di-GMP regulation.
Environmental
Carbon dioxide as the main greenhouse gas is going to be a global question. However, terrestrial plants could reduce only 30% of the CO2 emission. The novel reducing CO2 emission method is considered by the main countries around the world. Carbon fixation pathways have been achieved in engineered bacteria, which are fully autotrophic. We aim to design and implement efficient carbon fixation pathways in chassis cells and generate useful and economic products using synthetic biology and systems biology methods.